Public Bioinformatics Platform
By building a transparent, end-to-end automated platform, we can help address a key challenge facing science everywhere - the reproducibility crisis.
Genome scale research produces massive amounts of data that need to be processed into something useful. In the past, it required a huge investment in computers and infrastructure, which limited this kind of work to very large organisations. This is no longer true.
Modern computing practices and open standards based cloud computing are now accessible to anyone, anywhere on an affordable pay for what you use basis.

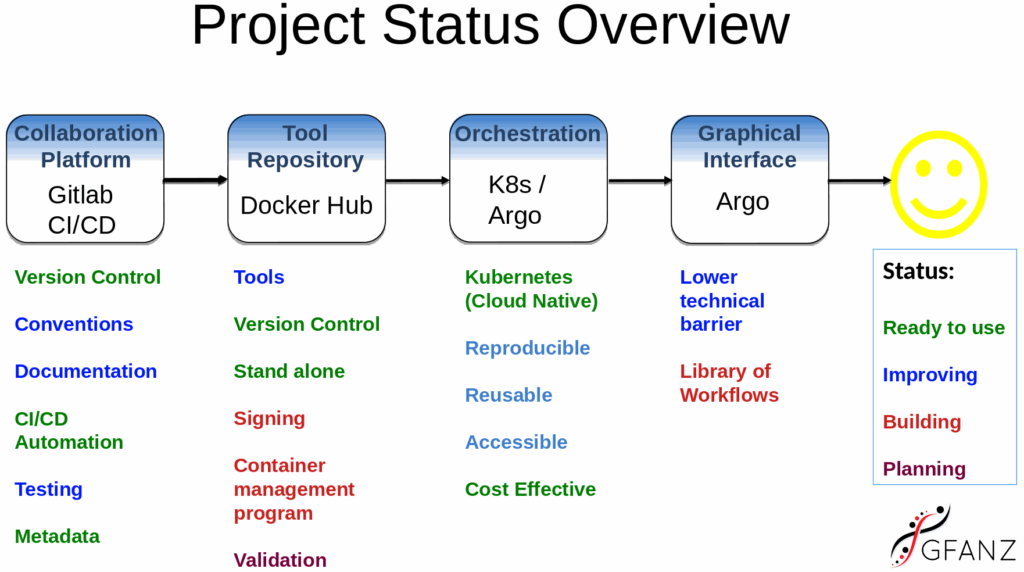
Platform Features
Through research about the state of bioinformatics tools and education done by the Palmerston North Bioinformatics Meeting group we defined the key set of issues in the field. We are addressing those issues by working with cutting edge open cloud computing technologies. Our aim is to build a best-of-breed bioinformatics platform that can be used by anyone, anywhere thereby democratising this area of research.
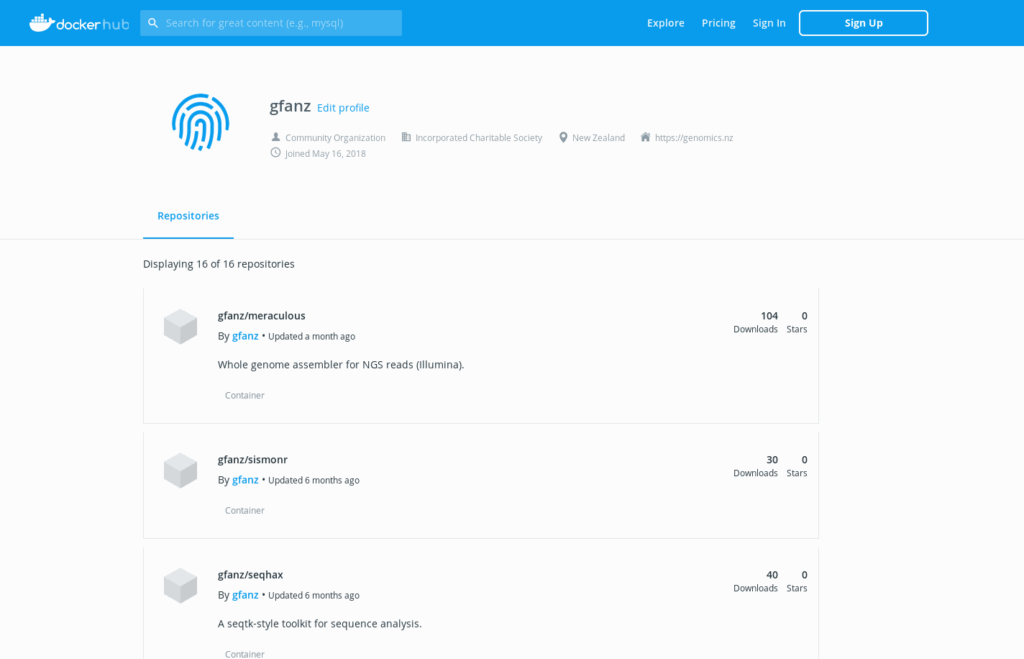
Software
There are many obstacles to using the best tool for the job in bioinformatics. This platform helps remove them so that the researcher can focus on using the right tool for their project.
Standards Based
The tools and platform are based on open standards, There is no vendor lock in.
All aspects are portable and sharable.
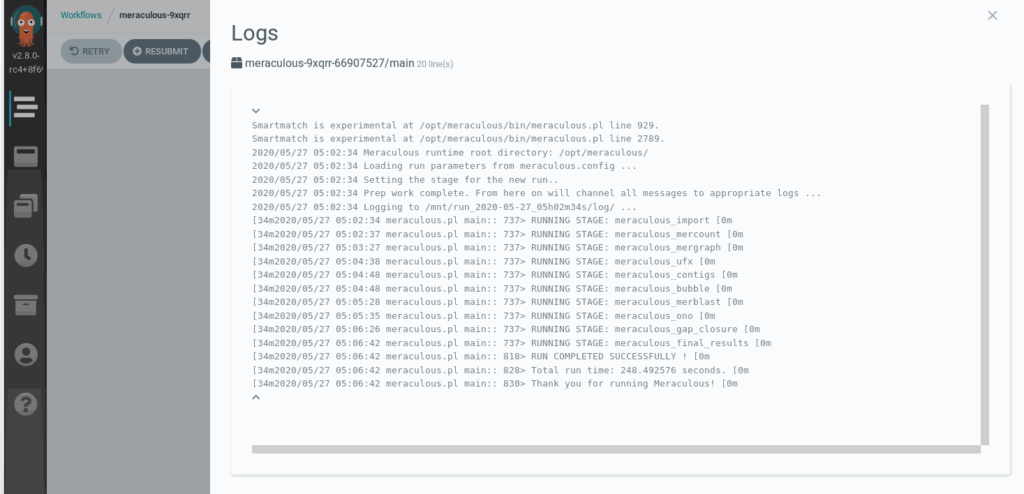
Workflows
We have chosen to use Argo as our workflow engine. It is a project of the Cloud Native Computing Computing Foundation. There are many big player supporting Argo. Using it, we can have a library reproducible workflows.
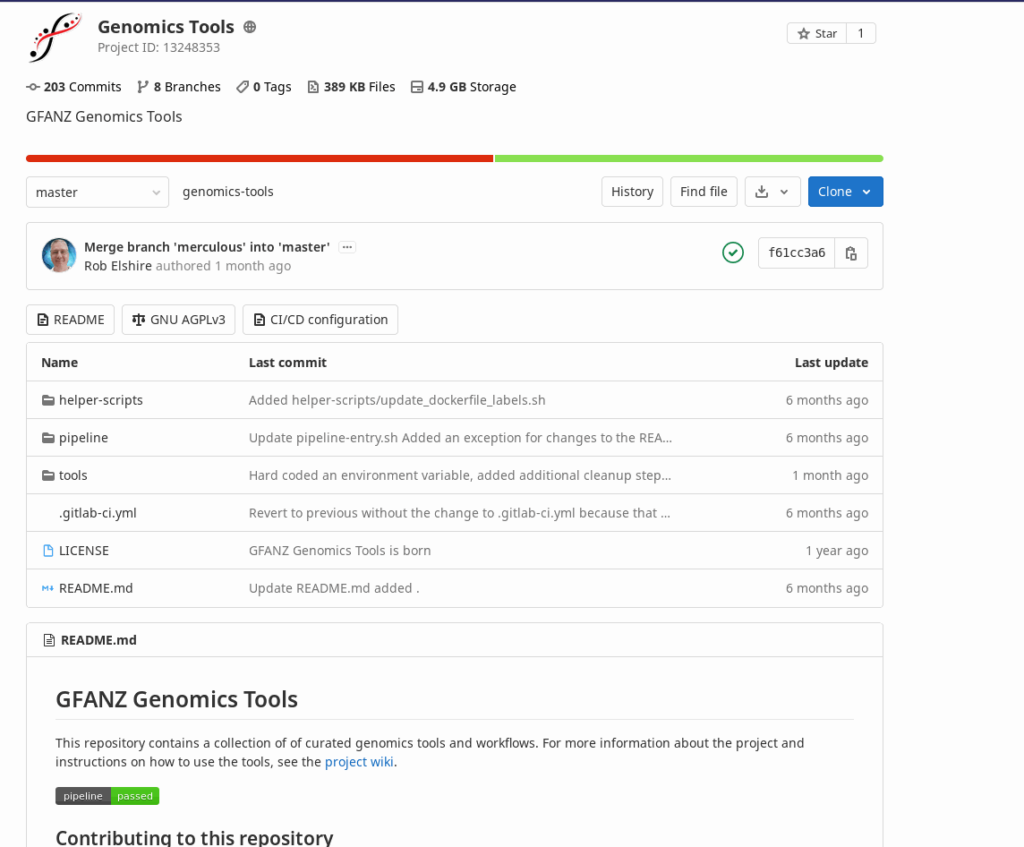
Collaboration
Our gitlab repository incorporates automation to test the scripts, and build and publish docker containers. All aspects of the project are under version control.
Anyone is welcome to participate.
Citations
Too often writers of bioinformatics software do not get credit for their work. This can hurt their careers. Our platform works to improve that situation by providing citation information for all the tools in a workflow.
Licensing
We want to ensure the work we do is available for anyone to use. Tools created using our collaboration are licensed under AGPL 3.0. Components used to create the tools must be licensed in a compatible way using any of these licenses.